Constrained-aware Diffusion for Scientific Applications
Constrained-aware Diffusion for Scientific Applications
Introduction
Generative AI, particularly denoising probabilistic diffusion models, has demonstrated remarkable success in synthesizing high-dimensional data. Their application is rapidly expanding beyond image synthesis into mechanical engineering, automation, chemistry, and biology.
However, a critical challenge limits their adoption in high-stakes scientific and engineering domains: while these models produce statistically plausible outputs, they often fail to adhere to physical principles, conservation laws, or safety constraints. Such violations result in suggested designs that may be impractical, mechanically unstable, or hazardous. Additionally, many scientific domains contend with sparse data and expensive data collection, demanding generalization to conditions never observed during training.
To address these challenges, our group has developed a class of training-free, constraint-aware generative models that integrate constraints and symbolic rules directly into the generative process through a novel combination of differentiable optimization and diffusion. The key idea is to reinterpret reverse generation, as a sequential optimization problem in which feasibility can be imposed during sampling, rather than checked only after the sample is complete. This creates a training-free interface between generative modeling and constrained optimization, and it opens the door to an application of differentiable constrained optimization directly inside the sampling loop.
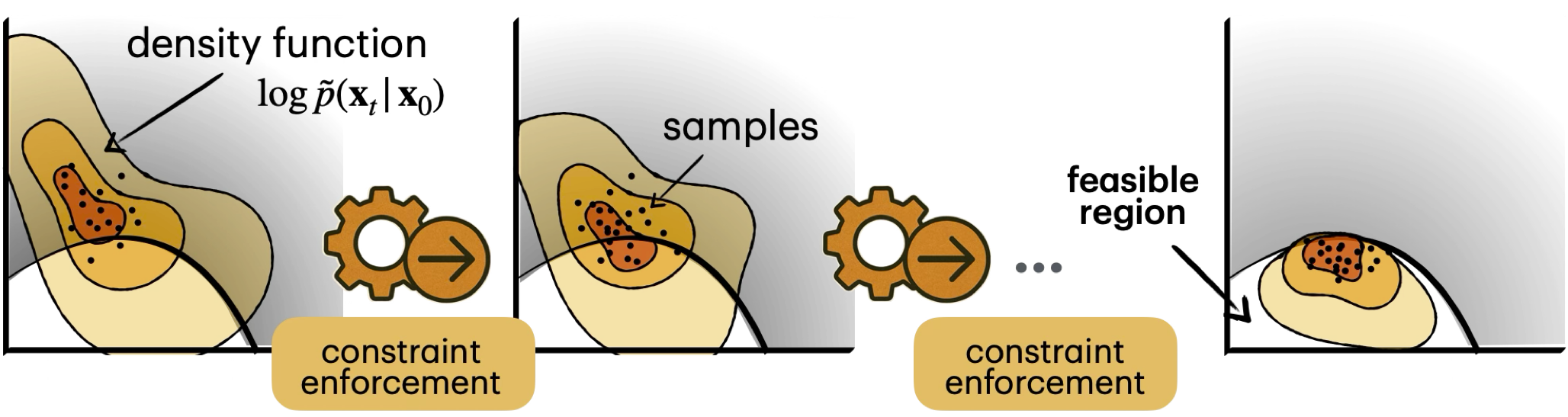
1. Foundational Methodology: Diffusion as Sequential Optimization
To understand our approach, we must first look at how standard diffusion operates. Denoising Diffusion Probabilistic Models define a forward stochastic process that progressively corrupts data $x_0 \sim p_{data}(x_0)$ into noise $x_T \sim p(x_T)$. This is often expressed as a stochastic differential equation (SDE) [cite]:
\[dx_t = -\frac{1}{2}\beta(t)x_t dt + \sqrt{\beta(t)}dB(t)\]where $\beta(t)$ defines an increasing noise schedule and $B(t)$ denotes a standard Brownian motion.
The reverse diffusion process recovers the original data via a time-reversed SDE. Since exact integration is intractable, it is discretized, and a neural network $s_\theta(x_t,t)$ is trained to approximate the score function $\nabla_{x_t}\log p(x_t)$. The reverse update is conventionally performed using Gradient Langevin Dynamics (GLD) [cite]:
\[x_t \leftarrow x_t + \gamma_t s_\theta(x_t,t) + \sqrt{2\gamma_t}\epsilon\]The Optimization Perspective and Projection
As the variance schedule decreases toward $t \rightarrow 0$, GLD transitions toward deterministic gradient ascent. This allows us to view the reverse diffusion process as a sequence of optimization problems minimizing the negative log-likelihood of the data distribution at each time step.
When application requirements impose constraints, we can frame the generation procedure formally:
\[\text{Minimize} -\log p(x_t|x_0) \qquad \text{Subject to:} \;\; x_T, \dots, x_0 \in C\]where $C$ defines the feasible region.
To account for these constraints, our Constrained-Aware Diffusion Models (CADM) introduce a projection operator [cite]:
\[\mathcal{P}_C(x_t) = \text{arg}\min_{y \in C} ||y - x_t||_2^2\]By applying this projection iteratively throughout the Markov chain, we take a gradient step to minimize the negative log-likelihood while ensuring feasibility. Theorem 1 of our work proves that for convex sets, the distance to the feasible set decreases at a rate of $1 - 2\beta\gamma_t$ at each step, reaching an $\epsilon$-feasible set in $\mathcal{O}(\gamma_{min}^{-1}\log(1/\epsilon))$ steps while preserving data fidelity.
Advanced Extensions: Lagrangians, Proximal Methods, and Flow Matching
Standard projections work well for simple convex geometries, but scientific constraints are often highly non-convex or hidden in latent spaces. We extend our framework through several advanced methods: Local Relaxations via Lagrangian Duality: For non-convex sets, we develop local relaxations based on the Augmented Lagrangian dual problem. By introducing multipliers $\lambda_j \ge 0$, we create an augmented projection objective that iteratively updates the sample and dual weights, allowing rapid inference without exact non-convex projections. * Latent Space Dynamics & Proximal Methods: Stable Diffusion operates in a low-dimensional latent space $z_t$, making input-space constraint evaluation difficult. We construct a computational graph that evaluates constraint violations $g(\mathcal{D}(z_t))$ post-decoding and propagates gradients back to the latent representation. We update $z_t$ using a proximal step [cite]:
\[\text{prox}_\lambda(g(\mathcal{D}(z_t))) = \text{arg}\min_{y} \left\{ g(\mathcal{D}(y)) + \frac{1}{2\lambda} ||\mathcal{D}(y) - \mathcal{D}(z_t)||_2^2 \right\}\]Flow-Matching Integration: We are extending CADM principles to flow-matching models, which transform a base noise distribution into a target distribution via a continuous-time velocity field $\frac{dx(t)}{dt} = v(x(t), t)$. By ensuring the velocity field keeps the sample trajectory within a specified feasible region, we can handle complex, probabilistic chance-constraints with highly efficient sampling. [cite]
2. Applications Across Scientific Domains
Our framework operates entirely during the sampling phase, acting as a training-free, plug-and-play module that preserves the underlying model’s inductive biases. We have deployed these techniques across four primary areas. [cite+1]
Protein Design with Hard Structural Constraints
Generating proteins requires strict adherence to geometric constraints (distances and angles) to ensure structural stability and substrate affinity.
Using RFdiffusion as our frozen backbone network, we enforce geometry and binding constraints natively during the generative loop. [cite]
By integrating iterative refinement, we scaffold novel protein backbones while preserving complex functional motifs, which are later validated using AlphaFold 3.
Materials Discovery and Simulators-in-the-Loop
Discovering structural materials often requires satisfying dynamic physical constraints, such as target porosity or specific stress-strain responses. [cite]
Since large-scale simulators (like Finite Element Analysis) typically lack differentiability, standard backpropagation fails.
We bypass this using sensitivity analysis, applying calibrated Monte Carlo perturbations to inputs to estimate pseudo-gradients of complex, non-smooth simulator outputs. This allows the diffusion model to adjust the material’s micro-structure dynamically until the simulated physical constraints are met. [cite]
Neuro-Symbolic Safety in Robotics and Control
Motion planning in autonomous systems requires finding collision-free paths governed by dynamic constraints, like ordinary differential equations (ODEs) for velocity and acceleration. [cite]
Our neuro-symbolic approach ensures that trajectories respect these physical boundaries and logic rules.
Remarkably, because we explicitly encode the physics, the model can generalize out-of-distribution (e.g., generating valid motion trajectories under lunar gravity despite only being trained on Earth’s gravity).
Constrained Discrete Diffusion for Chemistry and Drug Discovery
Chemical design is fundamentally discrete; generating molecules in SMILES format or as graphs requires selecting tokens from a finite vocabulary.
We adapted the CADM principle to discrete diffusion models by defining projections over the probability simplex that minimize the Kullback-Leibler divergence between the original logits and the updated, feasible logits [cite].
Because the $\text{argmax}$ operator is non-differentiable, we use Gumbel-Softmax relaxations to sample soft token probabilities.
This allows us to impose sequence-level constraints, such as BRENK substructure filters, ensuring near-perfect chemical validity while avoiding toxic substructures.
Conclusion
The future of generative AI lies not just in creating anything, but in creating exactly what we need. By embedding constraints natively into continuous diffusion, discrete diffusion, and flow matching frameworks, our group is bridging the gap between theoretical machine learning and applied science. Whether we are discovering life-saving drugs, designing novel materials, or ensuring robots operate safely, constrained generation is the key to unlocking the next era of reliable, scientifically grounded AI.
Interested in trying these models out? Code and pre-trained weights for the papers mentioned above are available on our lab’s GitHub page.